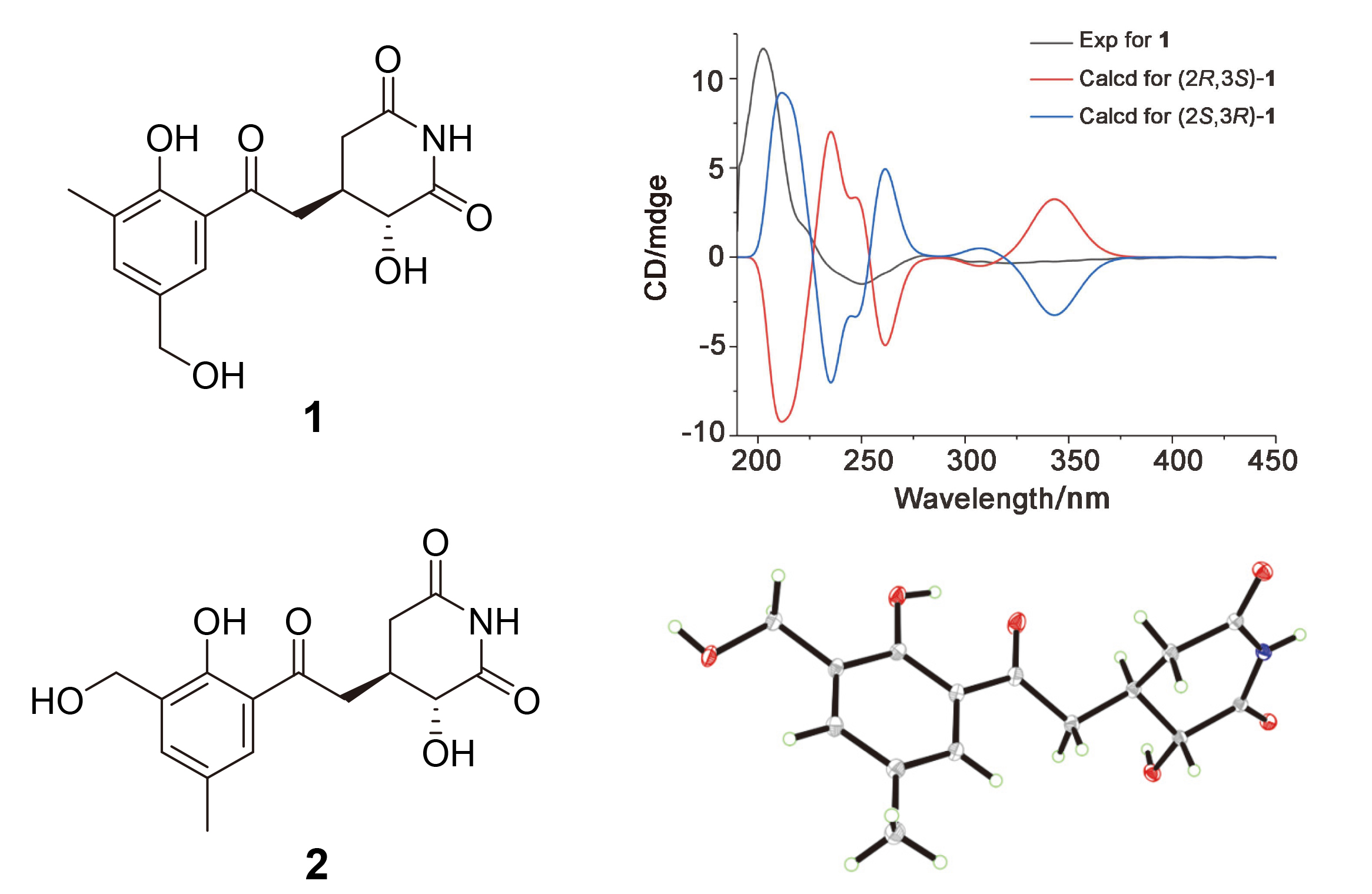
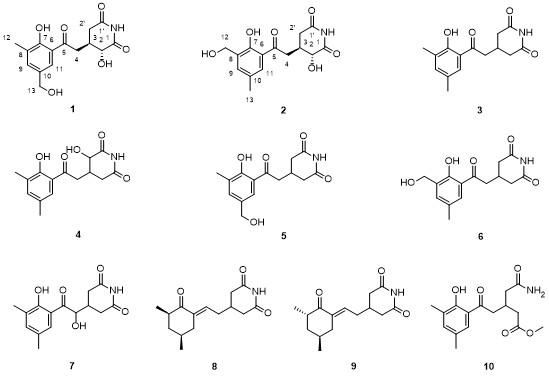
Compound
1 has an outward appearance of pale yellow solid. HR-ESI-MS analysis (
m/
z 306.0990 [M-H]
-, calcd for C
15H
16NO
6 306.0983) of
1 revealed its molecular formula as C
15H
17NO
6, containing eight degrees of unsaturation. Analysis of the
1H NMR spectrum (
Table 1) revealed the presence of one methyl, three methylenes, along with four methine protons. The
13C NMR and distortionless enhancement by polarization transfer (DEPT) indicated the presence of fifteen carbon resonances, which contained seven non-protonated carbons (
δC 206.5, 176.1, 173.8, 161.2, 132.8, 128.4, and 119.6), four methines (
δC 137.6, 127.5, 72.1, and 35.4), three methylenes (
δC 64.6, 41.0, and 37.4), and a methyl (
δC 15.5). Structural characterization of compound
1 was achieved through detailed NMR analysis and comparison with the known analog 3-[2-[2-hydroxy-3-methylphenyl-5-(hydroxymethyl)]-2-oxoethyl] glutarimide (
5).
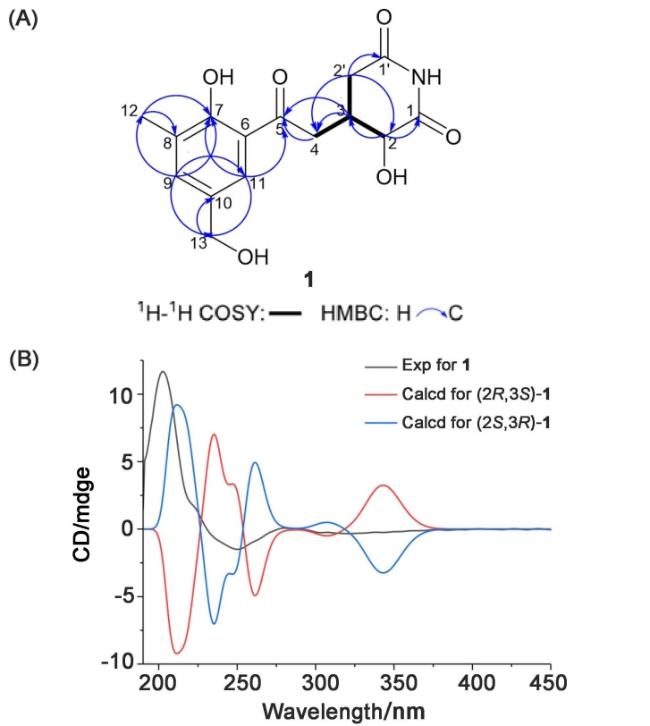
[5] Through systematic NMR analysis (
Figure 2A), it is first established that compound
1 shares the same carbon skeleton with
5 but features significant structural modification. Definitive heteronuclear multiple bond correlation spectroscopy (HMBC) correlations (H-9/C-7, C-11, C-12, C-13; H-11/C-5, C-7, C-9, C-13; H-12/C-7, C-8, C-9; H- 13/C-9, C-10, C-11) unambiguously confirmed the presence of a 1,2,3,5-tetrasubstituted benzene ring, while the continuous
1H-
1H correlation spectroscopy (COSY) coupling network [H-2 (H-2')/H-3/H-4] along with characteristic HMBC correlations (H-2/C-1, C-3, C-4; H2'/C-1', C-2, C-3, C-4; H-3/C-4, C-5; H-4/C-2, C-3, C-5; H-11/ C-5] collectively verified the glutarimide ring system fused with the benzene moiety. Particularly noteworthy was the significant downfield shift of C-2 (
δC 72.3, Δ+35.3) in compound
1 compared to
5. This observation, combined with the molecular formula analysis, conclusively demonstrated the hydroxyl substitution at C-2. Based on these multidimensional evidences, compound
1 was unequivocally identified as 2-hydroxy-3-[2-(2-hydroxy-3-methyl- phenyl-5-hydroxymethyl)-2-oxoethyl] glutarimide, representing a novel C-2 hydroxylated derivative of compound
5.