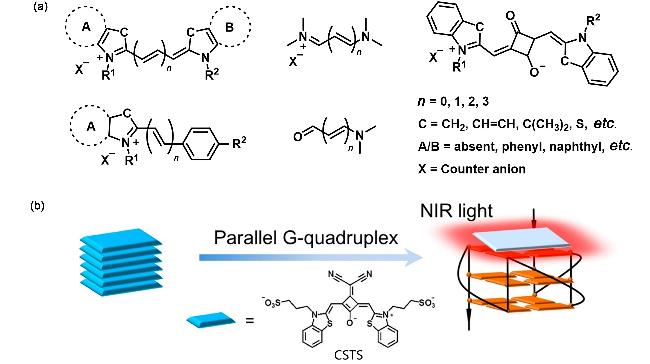
1 Introduction
2 G-quadruplexes (G4s)-response fluore- scence probes
2.1 Elastine dyes
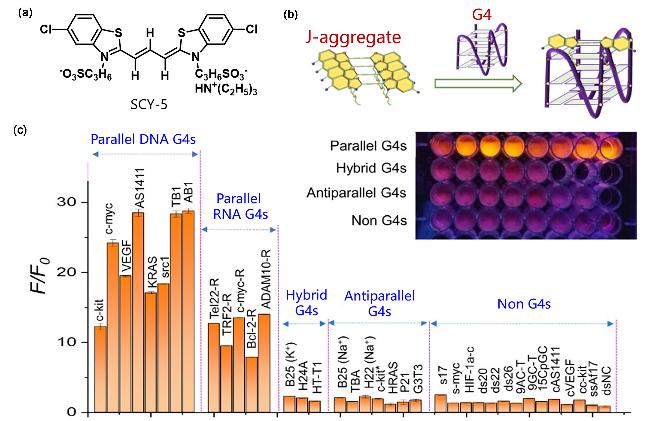
Figure 3 (a) Structural formula of SCY-5; (b) Luminescence phenomenon of SCY-5 inducing the disassembly of parallel G4s upon binding; (c) Fluorescence intensity (F/F0) of SCY-5 at 590 nm in the presence of different nucleic acid modelsReproduced from [44]. Copy right 2024 American Chemical Society. |
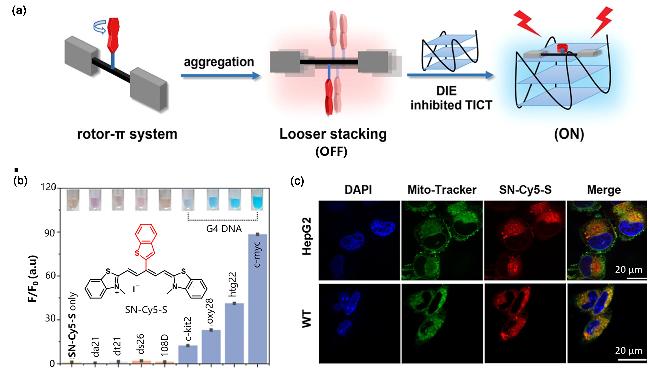
Figure 4 (a) Combination of SN-Cy5-S monomer with G4s inhibited twisted intramolecular charge transfer (TICT) and turn-on fluorescence; (b) Fluorescence enhancement (F/F0) of SN-Cy5-S after interaction with different nucleic acids; (c) Confocal fluorescence image of SN-Cy5-S mitochondrial co-localizationReproduced from [45]. Copy right 2023 American Chemical Society. |
2.2 Thiazole orange dyes
2.3 Thioflavin dyes
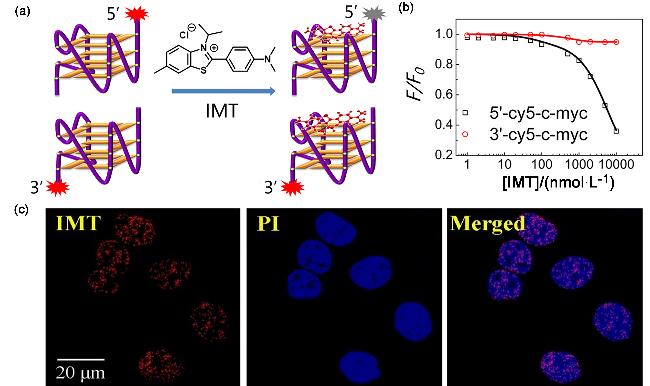
Figure 9 (a) Schematic diagram of the selective binding of IMT and G4s; (b) Relationship between F/F0 after the combination of IMT and c-kit and the concentration of IMT; (c) Confocal imaging of Hela cells stained and fixed with IMT and PIReproduced from [51]. Copy right 2018 Oxford University Press. |
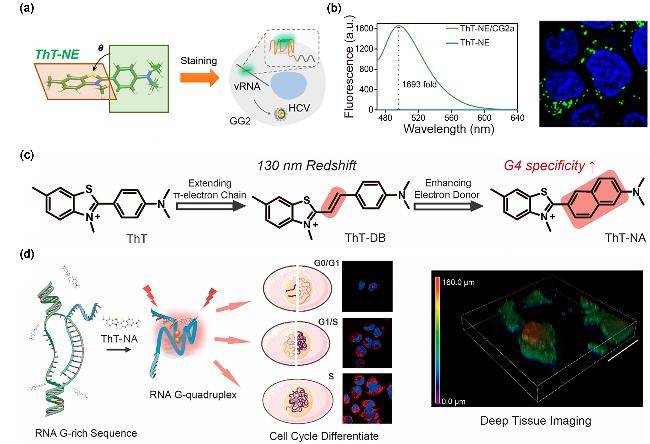
Figure 10 (a) Schematic diagram of the principle of detecting HCV vRNA using G4s probe ThT-NE; (b) Fluorescence spectra of ThT-NE in the presence of CG2a and direct intracellular visualization of CG2a by this probe; (c) Molecular structure, probe properties and modification of ThT; (d) Differentiation of cell cycle by ThT-NA through RNA G4 formation signal(a, b) Reproduced from [52], Copy right 2019 American Chemical Society.; (c, d) Reproduced from [53], Copy right 2022 American Chemical Society. |
2.4 Other small molecule dyes
2.5 Bioactive substance-based probes
2.6 Aggregation-induced emission probes
3 Development of fluorescence probes for other atypical nucleic acid structures
3.1 i-Motif sensing
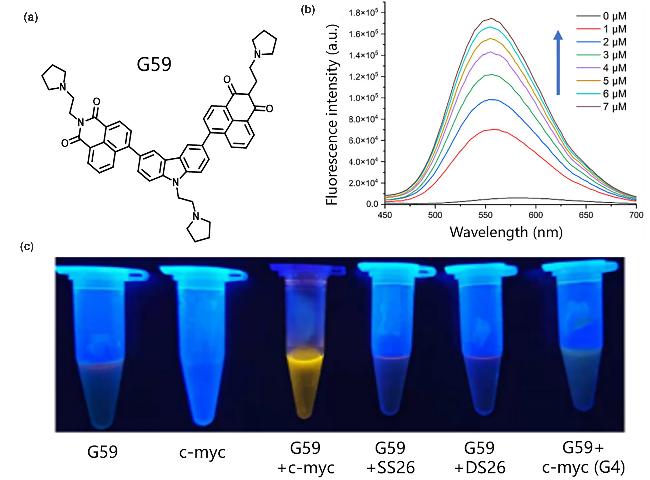
Figure 25 (a) Schematic diagram of the structure of G59; (b) Fluorescence emission spectra of G59 after adding different concentrations of i-motif; (c) Fluorescence changes of G59 binding with i-motif containing c-MYC promoter and other DNAReproduced from [94]. Copy right 2022 MDPI. |
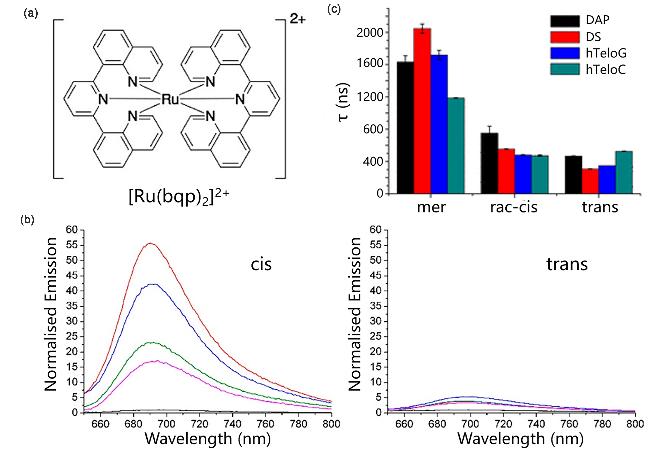
Figure 26 (a) Structurs of [Ru(bqp)2]2+; (b) Normalized emission intensities of cis,trans in the absence of DNA (black), hTeloC (green), DAP (blue), DS (red), and hTeloG (pink); (c) Fluorescence lifetimes of cis, trans and monomers under different conditionsReproduced from [95]. Copy right 2020 American Chemical Society. |
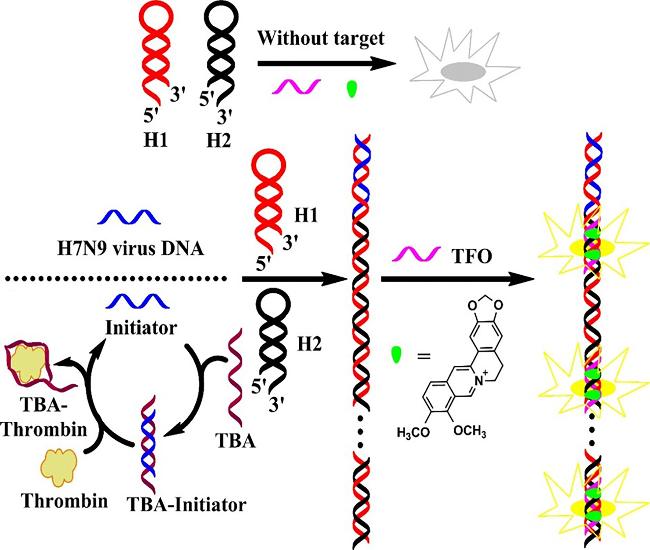
3.2 Triplex detection
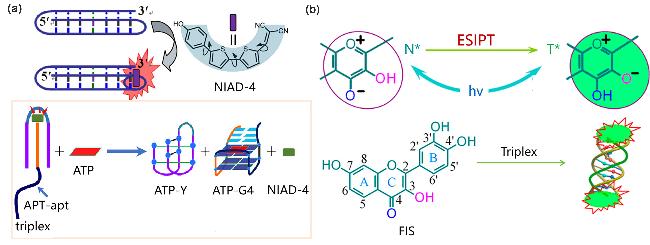
Figure 28 (a) Schematic diagram of Triplex detection mechanism of NIAD-4 and adenosine triphosphate (ATP) sensor based on Triplex; (b) FIS combined with Triplex to trigger ESIPT process to turn on fluorescence.(a) Reproduced from [100]. Copy right 2023 American Chemical Society; (b) Reproduced from [101]. Copy right 2015 American Chemical Society. |
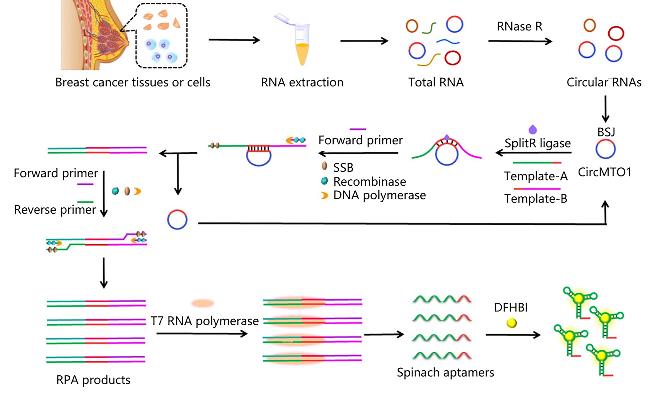
3.3 Circular nucleic acid (circDNA/RNA) analysis
4 Summary and prospects
Table 1 Comparison of different fluorescent probes for the detection of atypical nucleic acids |
| Probe | Types of nucleic acid | Key features | Ref. |
|---|---|---|---|
| CSTS | G4s | Disaggregation-Induced Emission (DIE), V-shaped rigid plane π bracket combined with the parallel G4s through the | [42] |
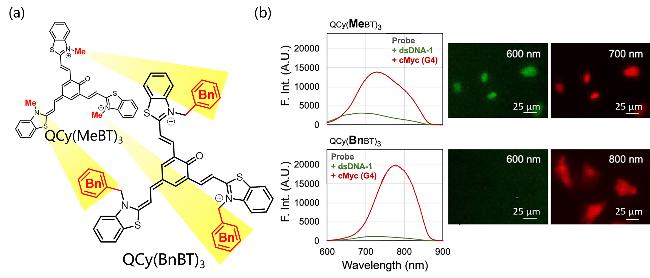
| QCy(BnBT)3 | G4s | High specificity recognition of G4s and 500 times fluorescence enhancement | [43] |
| SCY-5 | G4s | J-Aggregates dissociate into monomers, high specificity recognition of parallel G4s | [44] |
| SN-Cy5-S | G4s | Disaggregation-Induced Emission (DIE), Y-shaped plane is combined with the top plane at the 5' end of G4s | [45] |
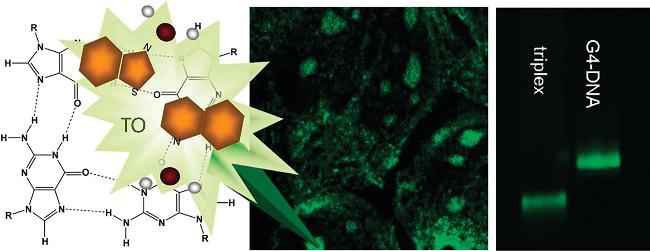
| TO | G4s and Triplex | Distinguish between Triplex and G4s, strong fluorescence enhancement | [47] |
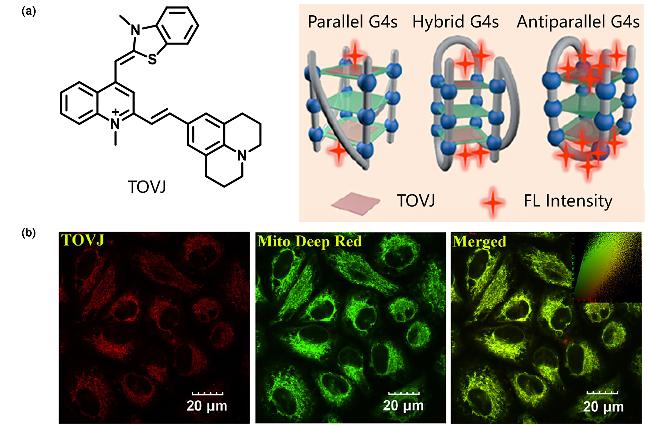
| TOVJ | G4s | High specificity recognition of antiparallel G4s, near-infrared emission | [48] |
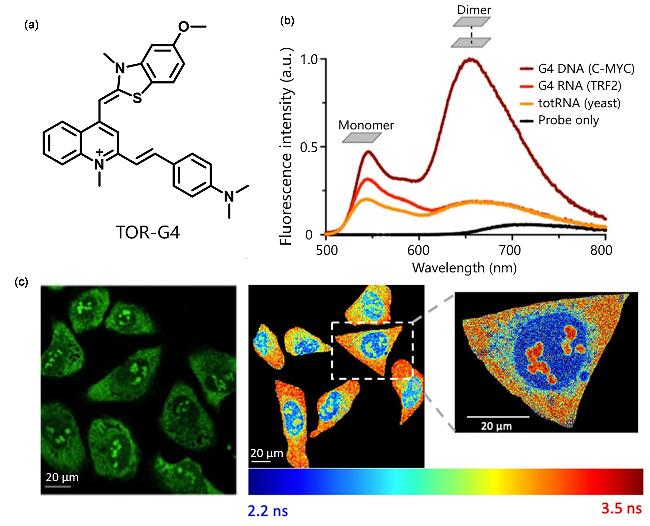
| TOR-G4 | G4s | Two-photon excitation, fluorescence lifetime has increased significantly | [49] |
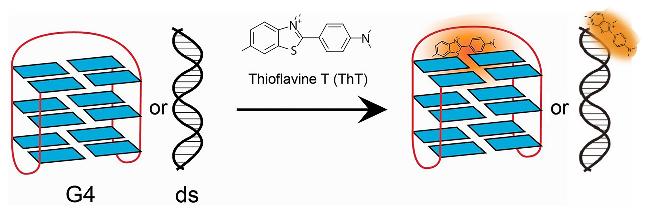
| ThT | G4s | High cell membrane permeability, specific recognition ability for G4s | [50] |
| IMT | G4s | Stack on the 3'-G4 end plane, low background and high signal-to-noise ratio emissions | [51] |
| ThT-NE | G4s | Intramolecular rotation is restricted and fluorescence is activated, high selectivity and sensitivity | [52] |
| ThT-NA | G4s | Red light emission, large Stokes shift, high fluorescence turn-on ratio and high selectivity for G4s | [53] |
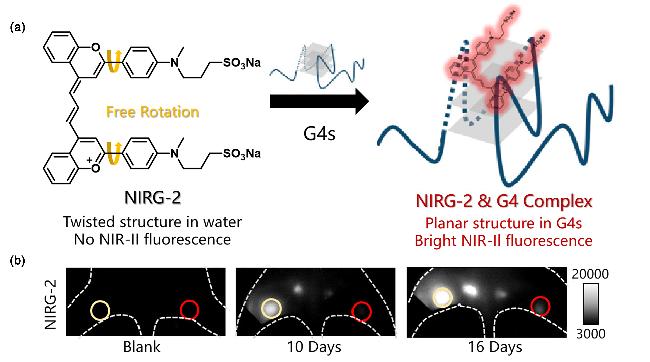
| NIRG-2 | G4s | Near-infrared Region II emission, hydrogen bonds and π-π stacking combine with G4s | [54] |
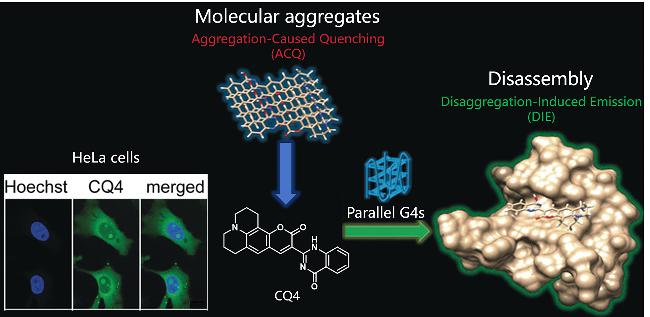
| CQ4 | G4s | Disaggregation-Induced Emission (DIE), specifically bind parallel G4s | [55] |
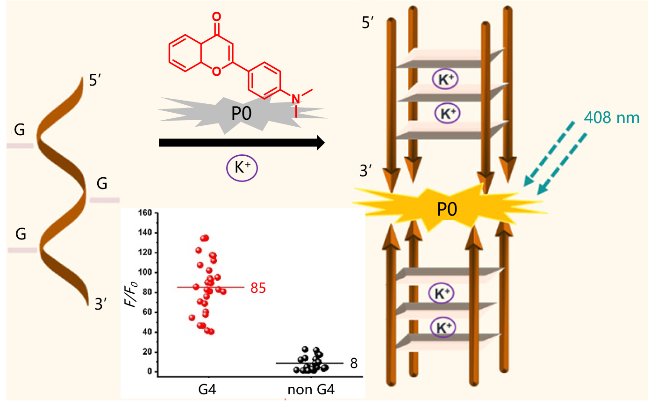
| P0 | G4s | The dimer G4/P0 system carried out highly selective detection of K+ | [56] |
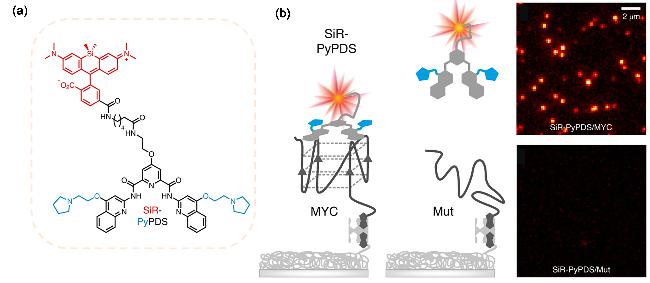
| SiR-PyPDS | G4s | The G4s ligand Pyridostatin (PyPDS) forms hydrogen bonds and hydrophobic interactions to specifically bind to G4s | [57] |
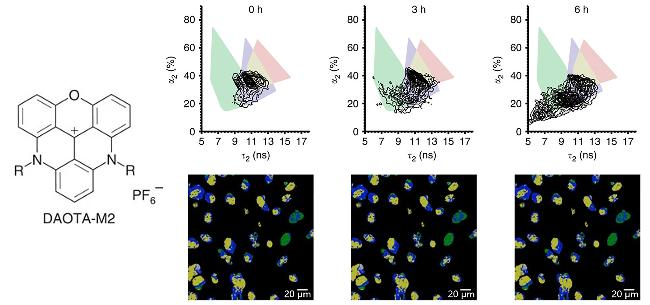
| DAOTA-M2 | G4s | G4s were identified by fluorescence lifetime imaging microscopy (FLIM) technology | [58] |
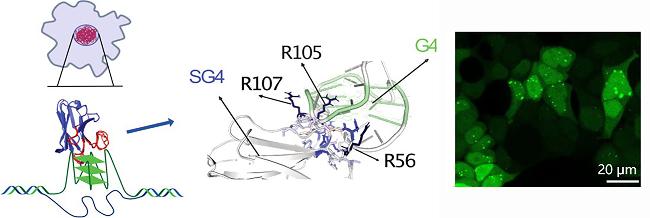
| SG4 | G4s | Antibodies against the structure of human c-MYC G4, bind to green fluorescent protein (GFP) | [59] |
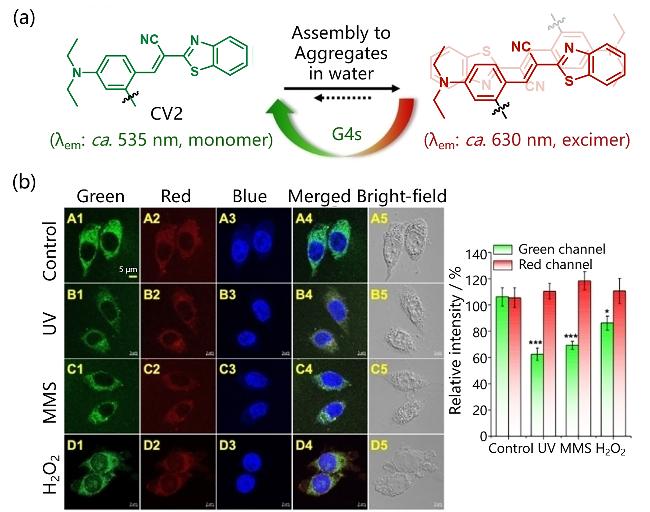
| CV2 | G4s | Peptide sequences specifically recognizing G4s (L-ARG-L-Gli-glutaric acid), Imaging of mitochondrial G4s | [60] |
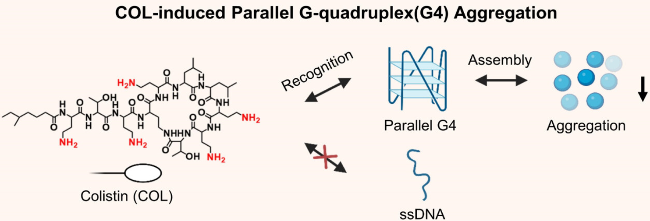
| COL | G4s | Induce parallel G4s to aggregate from the nucleic acid mixture | [61] |
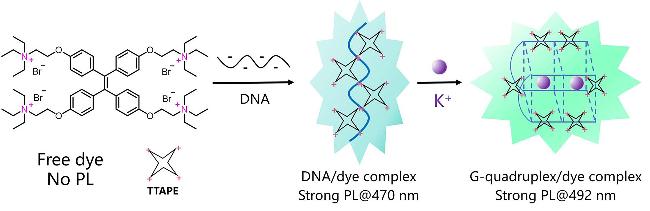
| TTAPE | G4s | Aggregation-induced emission detection (AIE), real-time monitoring of the folding process of G-rich DNA strands to form G4s | [84] |
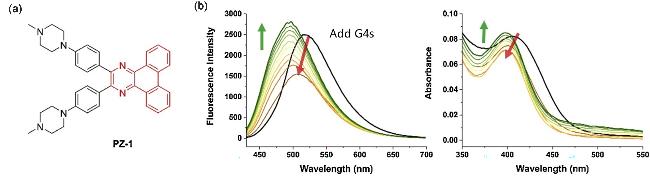
| PZ-1 | G4s | Aggregation-induced emission detection (AIE), G4s is combined through electrostatic interaction and π-π stacking | [85] |
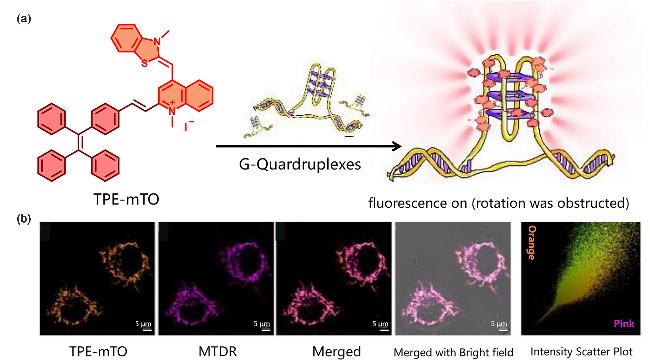
| TPE-mTO | G4s | Electron-deficient cation conjugated systems and crescent-shaped scaffolds, specifically locate G4s in mitochondria | [86] |
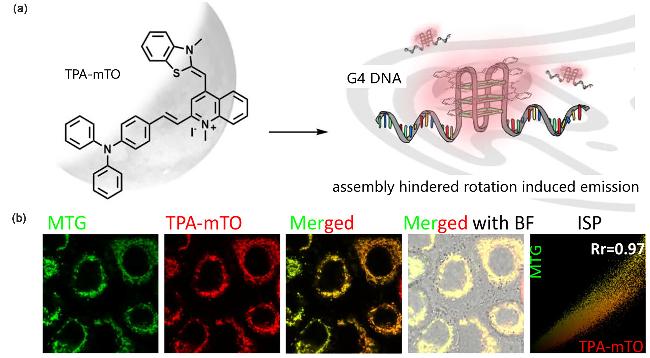
| TPA-mTO | G4s | Near-infrared emission, ideal photostability, and high fluorescence contras | [87] |
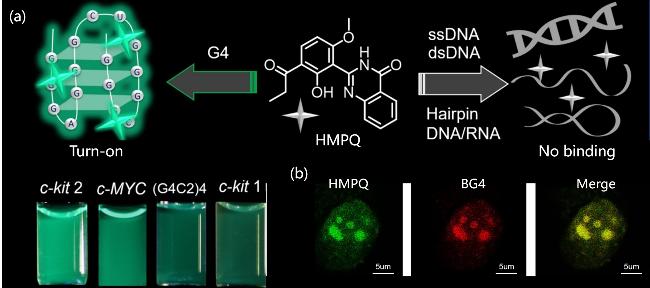
| HMPQ | G4s | AIEgens of biological origin, the π-π interaction binds to G4s, high selectivity and high sensitivity | [88] |
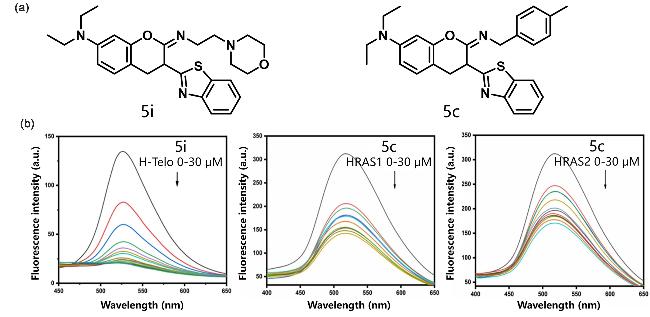
| 5i and 5c | i-motif | Combining different types of I-motifs, the fluorescence intensity is enhanced and the fluorescence lifetime is increased | [93] |
| G59 | i-motif | Specifically recognize and visually detect the i-motif structure of the c-MYC gene promoter | [94] |
| [Ru(bqp)2]2+ | i-motif | The cis isomer combines with the DAP i-motif, resulting in fluorescence activation and increased fluorescence lifetime | [95] |
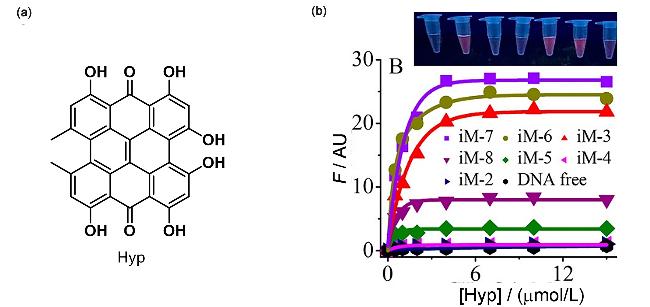
| Hyp | i-motif | The i-motif structures of different lengths were distinguished by fluorescence intensity | [97] |
| NIAD-4 | Triplex | Uncharged near-infrared molecular rotor probe, topological match with the Triplex terminal triad | [100] |
| FIS | Triplex | The quantity ratio of 2∶1 is combined with Triplex, the ESIPT process is triggered and the green fluorescence lights up | [101] |
| Berberine | Triplex | Isoquinoline alkaloids, combined with the Triplex structure, it will show strong fluorescence at 530 nm | [102] |
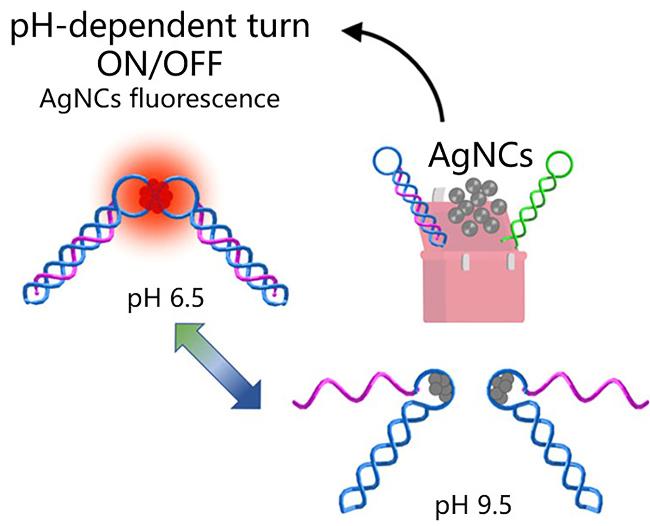
| AgNCs | Triplex | The dynamic changes of photophysical properties with the Triplex structure | [103] |
| DFHBI | circRNA | Using circMTO1 as the template, the aptamers generated by RPA and transcriptional amplification techniques were combined with DFHBI | [104] |
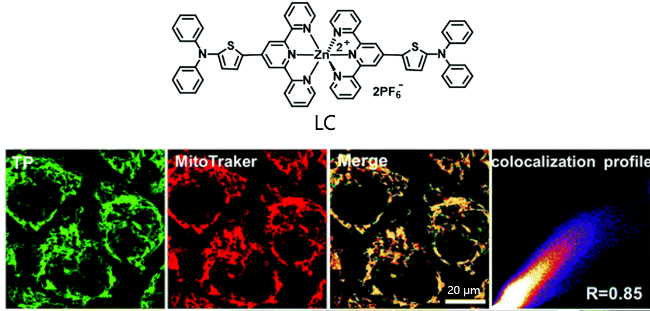
| LC | circDNA | Efficient intracellular uptake capacity, high-resolution visual imaging detection of mitochondrial DNA (mtDNA) | [106] |
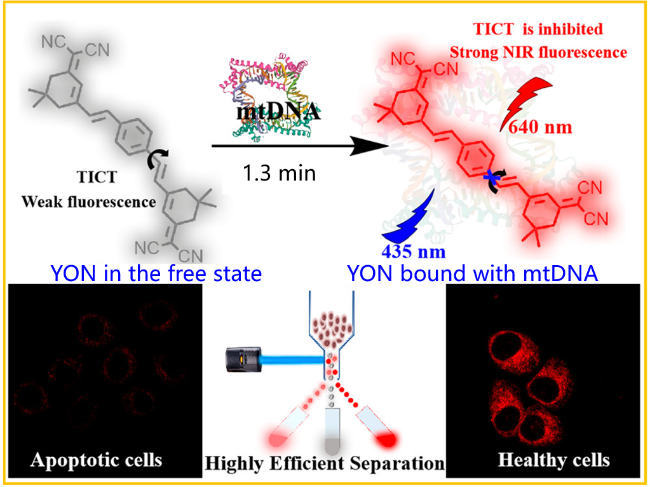
| YON | circDNA | Near-infrared twisted intramolecular charge transfer (TICT) fluorescent probe, high sensitivity and large Stokes displacement | [107] |