Compound
1 was obtained as a white amorphous powder. Its molecular ion peak was determined by HRESIMS at
m/
z 264.1594 [M+H]
+ (calcd for C
15H
22NO
3 264.1600), corresponding to a molecular formula C
15H
21NO
3 with six degrees of unsaturation. The ¹H NMR spectrum displayed characteristic signals including two singlet methyl groups at
δH 1.06 (s, 3H) and 1.07 (s, 3H), one doublet methyl group at
δH 1.38 (d,
J=6.4 Hz, 3H), and one olefinic proton at
δH 6.50~6.52 (m, 1H). The ¹³C NMR spectrum showed 15 distinct carbon signals, featuring: one ester carbonyls (
δC 176.05), one amide carbonyl (
δC 169.59), two olefinic carbons (
δC 140.15 and 143.94), and one oxygenated methine (
δC 80.18) (
Table 1). The
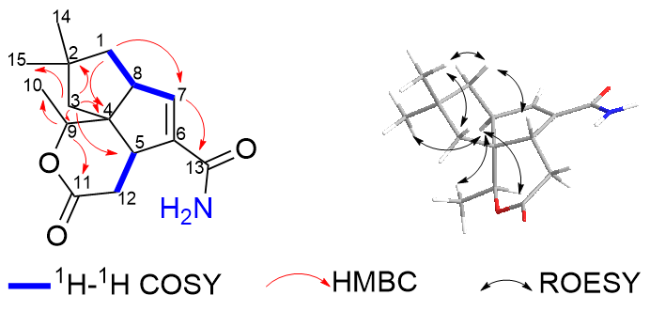
1H-
1H correlation spectroscopy (COSY) spectrum indicated the presence of three isolated proton sequences of H
2-1/H-8/H-7, H-9/H
3-10 and H-5/ H
2-12 (
Figure 2). The above NMR data suggests that
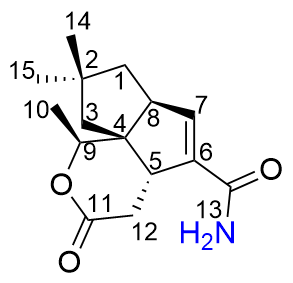
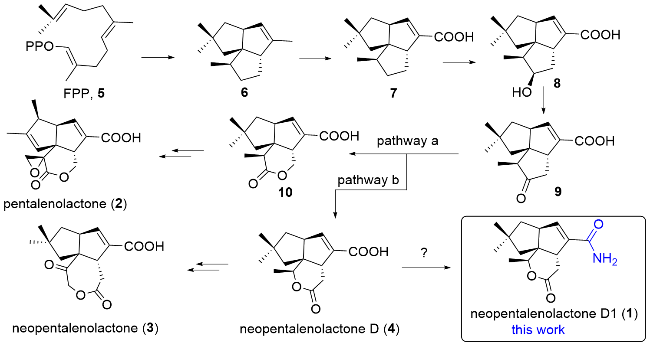
1 is a sesquiterpene structurally analogous to neopentalenolactone D (previously identified as its methyl ester
[10-11]) except for the major chemical shift differences observed at C-13 (
δ +2.23), C-6 (
δ +2.51), and C-7 (
δ -5.71). The heteronuclear multiple bond correlations (HMBC) observed between H-9 to C-10 and C-11, and H-1 to C-7, and H-7 to C-13 provided further confirmation that
1 shares the same molecular skeleton with neopentalenolactone D. In combine with the molecular formula, the number of hydrogens directly bonded to carbon and the index of unsaturation, there is an NH
2 group to be assigned. Based on the molecular scaffold established by 1D and 2D NMR, it is reasonable that the extra NH
2 group was linked to C-13 carbonyl, which was confirmed by the chemical shift differences at C-13, C-6, C-7, thus the chemical structure of compound
1 was established as the amide derivative of neopentalenolactone D (
Figure 2).