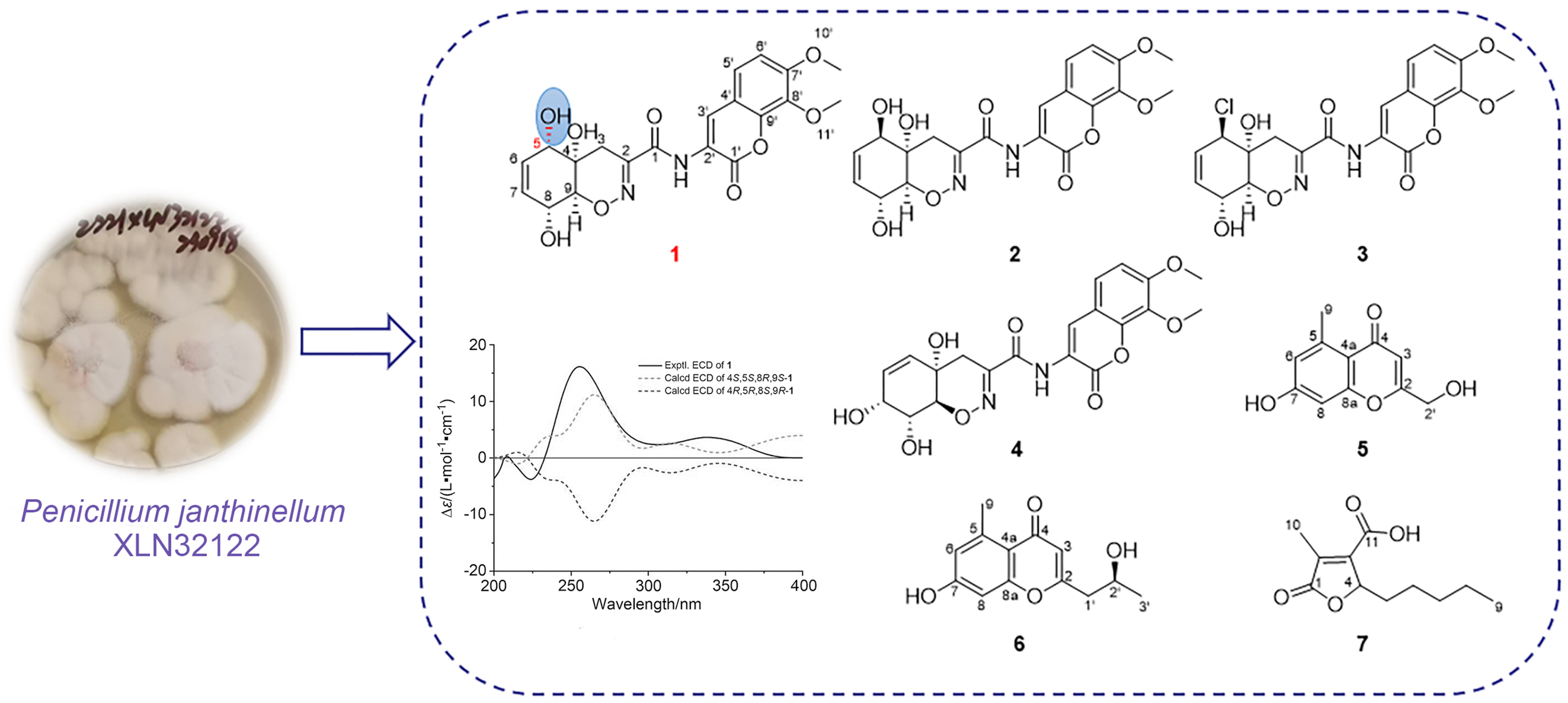
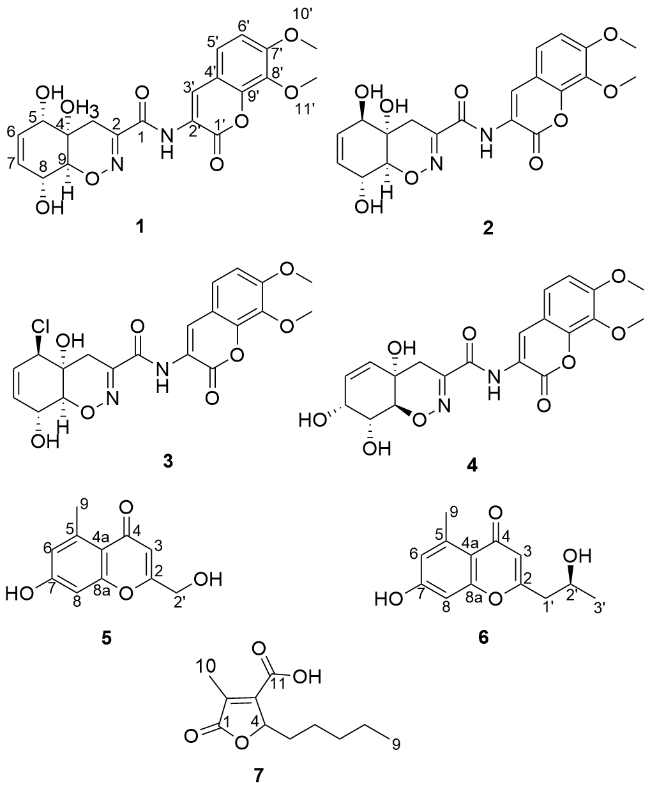
Trichodermamide H (
1) was isolated as a yellow amorphous powder. The molecular formula of compound
1 was established as C
20H
20N
2O
9 based on HR-ESI-MS [M+ Na]
+ ion peak (calcd for C
20H
20N
2O
9Na 455.1066, found 455.1056), indicating 12 degrees of unsaturation. The
1H NMR data (
Table 1), combined with the heteronuclear multiple quantum coherence (HMQC) spectrum of
1 in DMSO-
d6, showed an exchangeable hydrogen signal at
δH 9.32 (s, 1H, NH), three hydroxyl groups at
δH 5.46 (d,
J=6.4 Hz,1H, 8-OH), 5.31 (d,
J=4.8 Hz, 1H, 5-OH), and 5.09 (s, 1H, 4-OH), two aromatic protons at
δH 7.49 (d,
J=8.8 Hz, 1H, H-5') and 7.13 (d,
J=8.8 Hz, 1H, H-6'), three olefinic protons at 5.69 (ddd,
J=10.4, 4.8, 2.4 Hz, 1H, H-6) and 5.48 (dd,
J=10.4, 2.4 Hz, 1H, H-7), three methine groups at
δH 3.82 (br d,
J=4.8 Hz, 1H, H-5), 3.97 (t,
J=7.6 Hz, 1H, H-8) and 4.11 (dd,
J=7.6, 2.0 Hz, 1H, H-9), two methoxy groups at
δH 3.89 (1H, s, H-10') and 3.83 (1H, s, H-11'), and one methylene group at
δH 2.07 (d,
J=18.8 Hz, 1H, H-3
α) and 2.28 (dd,
J=18.8, 2.4 Hz, 1H, H-3
β). The
13C NMR data (
Table 1), combined with distortionless enhancement by polarization transfer (DEPT)-135 spectrum of
1, showed the presence of two carbonyl carbons at
δC 160.9 (C-1) and 157.9 (C-1'), 11 aromatic and olefinic sp
2-carbons at
δC 149.9 (C-2), 127.4 (C-6), 130.5 (C-7), 121.0 (C-2'), 123.9 (C-3'), 1113.7 (C-4'), 123.1 (C-5'), 110.2 (C-6'), 53.9 (C-7'), 135.3 (C-8') and 143.6 (C-9'), three oxygenated methine groups at
δC 69.4 (C-5), 66.4 (C-8) and 63.9 (C-4), two methoxy carbons at
δC 56.5 (C-10') and 60.9 (C-11'), one oxygenated quaternary sp
3-carbon at
δC 81.7 (C-9), and one methylene carbon at
δC 26.3 (C-3). The
1H NMR and
13C NMR data of compound
1 were similar to those of trichodermamide A (
2),
[28] and had the same molecular formula as
2 on the basis of the HR-ESI-MS data. The major differences were the chemical shifts of C-5 at
δH 3.82 (br d,
J=4.8 Hz, 1H) and
δC 69.4 (CH) in
1, instead of at
δH 4.24 (br d,
J=2.4 Hz, 1H) and
δC 73.0 (CH) in
2, and the chemical shifts of C-9 at
δH 4.11 (dd,
J=7.6, 2.0 Hz, 1H) and
δC 81.7 (CH) in
1, in contract to the chemical shifts at
δH 3.99 (br s, 1H) and
δC 83.8 (CH) in
2. The above results indicated that compounds
1 and
2 had the same plane structure, but differ in their configurations at the cyclohexane ring. The relative configuration of
1 was determined on the basis of nuclear overhauser effect spectroscopy (NOESY) and coupling constants. The NOESY correlations of H-5 to H-8 indicated that H-5 and H-8 were on the same side of the cyclohexane ring, and the correlations of 4-OH to 5-OH/8-OH, and 5-OH to 8-OH suggested that 4-OH, 5-OH and 8-OH oriented to the other side of the cyclohexane ring (
Figure 2). In addition, the large coupling constants between H-8 (7.6 Hz) and H-9 (7.6, 2.0 Hz), consistent with literature data,
[14,28] further confirmed their
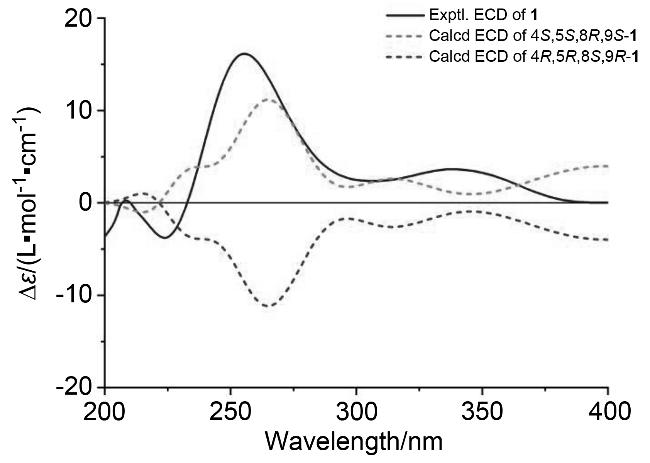
trans-diaxial orientation. The absolute configuration of
1 was determined on quantum chemical electronic circular dichroism (ECD) calculations. The theoretical ECD spectra of two possible stereoisomers of 4
S,5
S,8
R,9
S and 4
R,5
R,8
S,9
R were created by the time-dependent density functional theory (TDDFT) calculations, and the calculated ECD spectrum of the isomer 4
S,5
S,8
R,9
S matched well with the experimental ECD curve of
1 (
Figure 3). Thus the structure of
1 was determined, and named as trichodermamide H (
1).



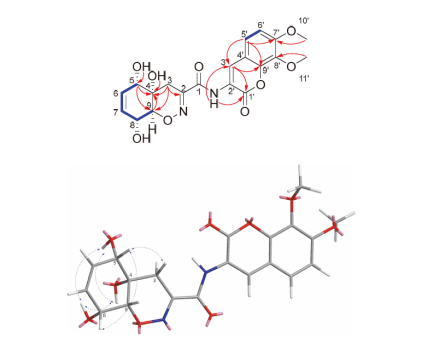
 ), HMBC (
), HMBC ( ) and NOESY (
) and NOESY ( ) correlations of compound 1
) correlations of compound 1